Disentangled representations
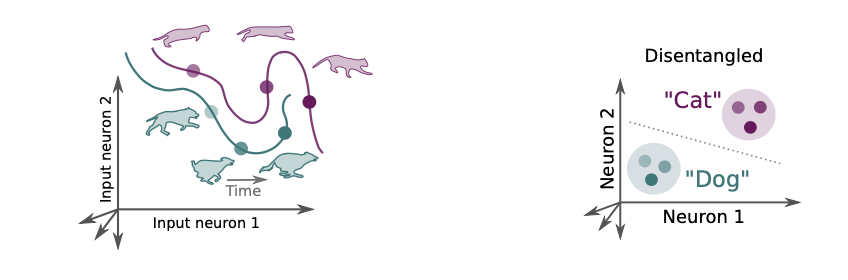
What is disentanglement exactly? How do we measure it?
The linear classification measure is limited to the classes we have labels for. It can be non-trivial to measure intuitive quantities. There are at least 3 notions of disentanglement out there:
- Each neuron codes for only one latent
- Each latent coded only by one neuron
- Representation trajectories are straightened?
A good review of the difficulties in this process is available at Locatello et al. (undefined) - A Sober Look at the Unsupervised Learning of Disentangled Representations and their Evaluation
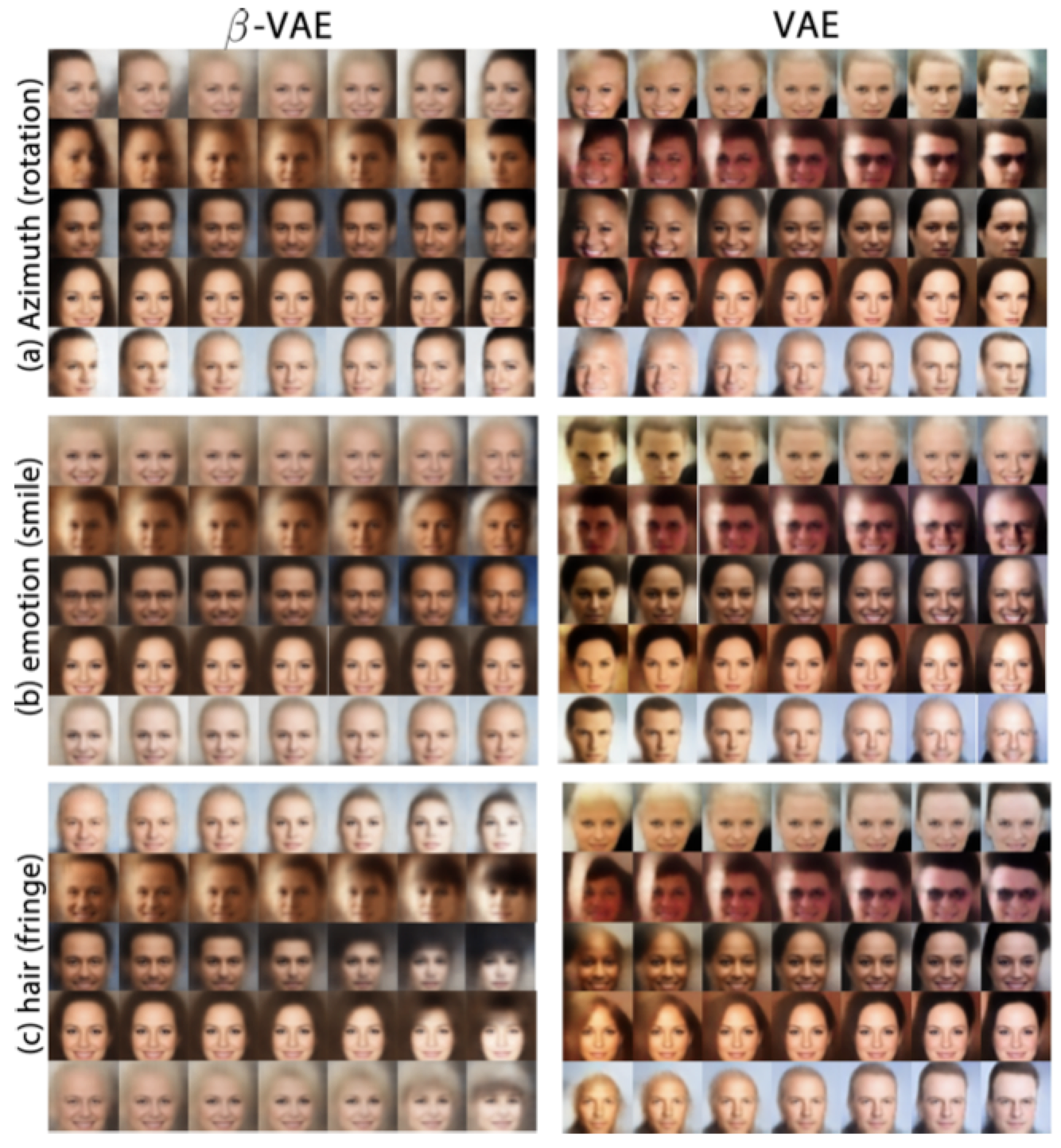
Qualitative analysis with VAEs
Strategy: vary one latent variable (in latent space) and observe whether changes in the output are limited to a single interpretable feature and that we do not end up interfering with the other features in the output (ex: only face angle or only facial expression).

Metric 1
Rationale: disentangled representations are both interpretable and independent.
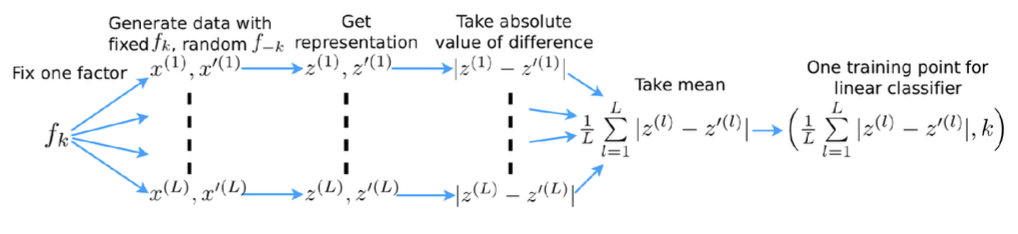
Problems:
- Depends on hyper parameters of the linear classifier
- Not axis-aligned - each factor could be represented by a linear combination of neurons instead of a single one
- Perfect score even when only K-1 out of K factors have been disentangled
Metric 2
Disentangled representations are both interpretable and independent
also axis-aligned, with fewer hyperparameters.
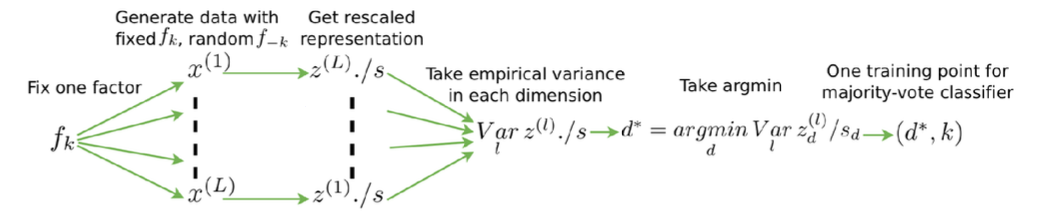
Problems:
- Classification-based metrics are ad-hoc and sensitive to hyperparameters
- Loosely speaking, they measure reduction in entropy of unit activities
given fixed values of latent factors
Mutual Information Gap (MIG)
Rationale: directly measure mutual information between representations and unit activity distributions.
MIG: Normalise with entropy, and more importantly, enforce axis-alignment by measuring the difference between the top-two neurons for a given factor
Problems:
- Limits the number of neurons per latent factor to exactly one
- Particularly bad when latent factor structure itself is 2D
Mutual Information Ratio (MIR)
Rationale: Disentangling is when single neurons care about single ground truth factors. We don’t mind if more than one neuron cares about the same factor.
Each neuron's preference:
MIR:
How to use MI measures for a deterministic network
- Discretize every neuron's activity in terms of bins defined on each neuron's activities across all data and calculate the MI between this discretized representation and the factor values.
- Basically compute MI between the discretized (independently per-neuron) empirical distribution of neuron activities and the corresponding empirical factor distribution. and since everything is discrete here, MI can be estimated unlike before discretization.
All the above measures give high scores to pixels because they do not normalize wrt total number of available dimensions.
MI-based Entropy Disentanglement score (MED)
Rationale: normalize the mutual information by columns, such that an entry in the matrix indicates the relative importance of one dimension over all dimensions regarding a certain data factor.
Importance matrix:
After normalizing over the columns, evaluate the contribution of a dimension to different factors, which is described by a row of
Disentanglement score
where
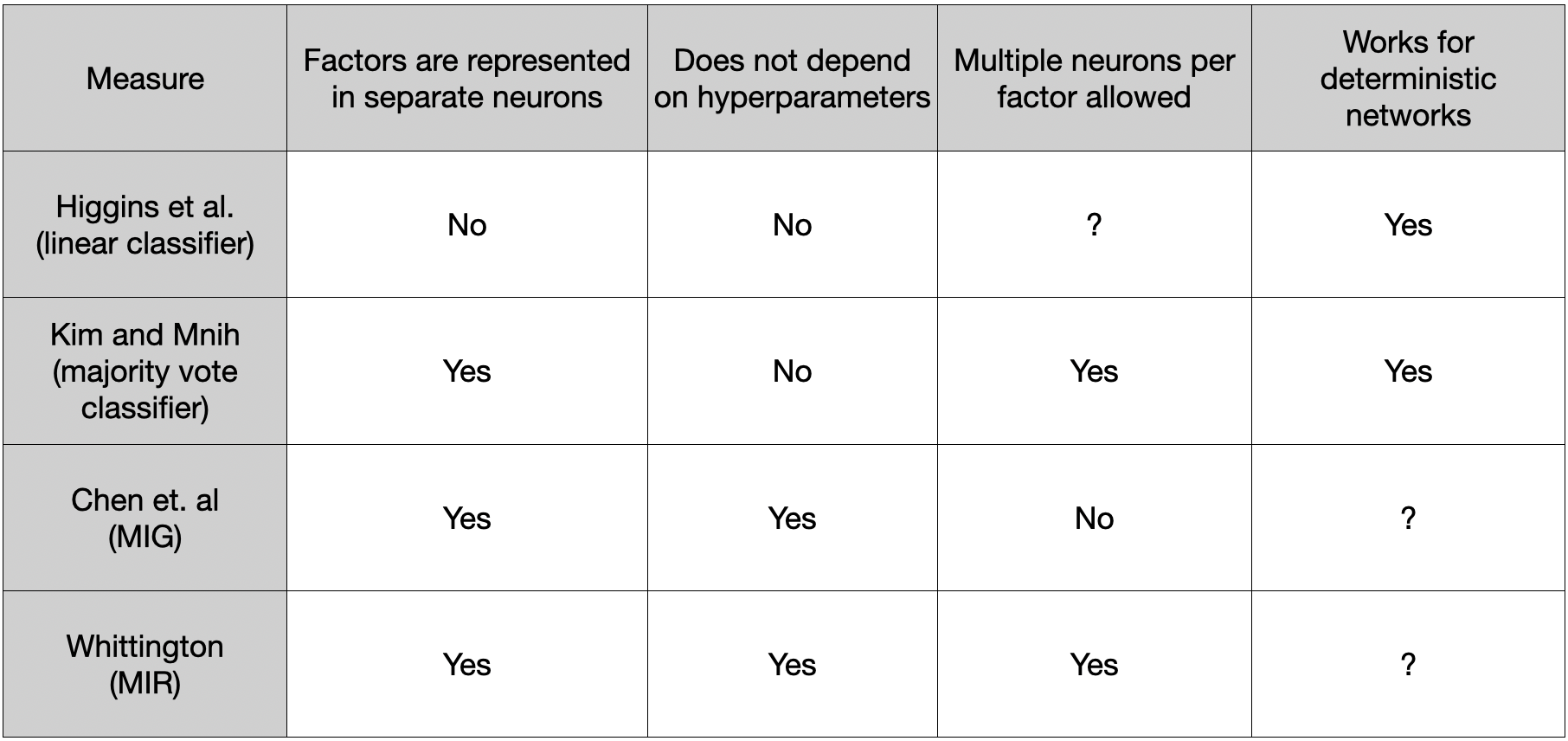
- Kim and Mnih (2019) - Disentangling by Factorising
- Burgess et al. (2018) - Understanding disentangling in
-VAE - Whittington et al. (2022) - Disentangling with Biological Constraints A Theory of Functional Cell Types
- Cao et al. (2022) - An Empirical Study on Disentanglement of Negative-free Contrastive Learning